OLake PoC: Real-Time PostgreSQL to Apache Iceberg CDC Without Kafka
This document summarizes a PoC process of setting up OLake — a lightweight tool that performs CDC directly from PostgreSQL to Apache Iceberg without Kafka — in a local Docker environment, and testing its performance, stability, and ease of use in comparison with Debezium.
Introduction
In a previous blog post, we built a CDC pipeline in the PAASUP DIP environment that synchronizes change data from PostgreSQL to StarRocks using Kafka and the Debezium Source Connector. While Debezium is a proven CDC tool, it comes with the operational overhead of managing supplementary infrastructure such as a Kafka cluster and Schema Registry.
Meanwhile, as data lakehouse architectures have gained traction, demand for CDC pipelines targeting Apache Iceberg has grown. For such use cases, Debezium requires an additional path via a Kafka → Iceberg Sink Connector, whereas OLake is a "Lakehouse First" lightweight CDC tool that pushes data directly from the source database to Iceberg — without Kafka.
In this blog post, we set up an OLake-based PostgreSQL → Apache Iceberg CDC pipeline in a local Docker environment and document the process of testing it for performance, stability, and ease of use. This test is part of a PoC to determine whether OLake should be included in the DIP catalog.
Table of Contents
- Debezium vs. OLake
- OLake Architecture
- Infrastructure Setup
- Loading Test Data into source_postgresql
- Configuring the OLake CDC Pipeline
- Querying Iceberg Tables
- OLake CDC Metadata Structure
- CDC Columns in Iceberg Tables
- Test Results
- Conclusion
1. Debezium vs. OLake
Debezium, used in a previous blog post, and OLake, tested this time, are both CDC tools — but their design philosophies and architectures are fundamentally different.
Design Philosophy
- OLake — Takes a "Lakehouse First" approach. It is optimized for pushing data as fast as possible directly from a source database to a data lakehouse such as Apache Iceberg, without the need for a complex Kafka infrastructure.
- Debezium — Takes an "Event-Driven" approach. It treats database changes as "events" and streams them to Kafka, enabling various downstream systems (search engines, caches, other databases, etc.) to subscribe to those events. It is a general-purpose CDC framework.
Architecture Comparison
| Feature | OLake | Debezium |
|---|---|---|
| Core Structure | Single binary/container-based. Connects directly from the source DB to the destination without a separate message queue. | Kafka ecosystem-based (requires Kafka, Zookeeper, and Kafka Connect). |
| Data Flow | DB → OLake → S3 (Iceberg/Parquet) | DB → Debezium → Kafka → Sink Connector → Destination |
| State Management | Stores checkpoints in local or designated storage. | Stores state in internal Kafka topics (offsets, config, status). |
| Scaling | Horizontal scaling is possible, but the structure is simpler. | Distributed processing via Kafka Connect cluster worker node scaling. |
Pros and Cons
| Category | OLake | Debezium |
|---|---|---|
| Pros | - 4–10x faster for bulk data loads by bypassing Kafka. | - Can deliver data to virtually any destination, not just lakehouses. |
| - No Zookeeper/Kafka management required, reducing operational burden. | - Familiar to organizations already using Kafka, with strong community support. | |
| - Automatic support for schema evolution and Iceberg optimization. | - Per-message transformation via SMT (Single Message Transforms). | |
| - Lower infrastructure resource usage, reducing cloud costs. | - Widely referenced across many enterprise environments. | |
| Cons | - Currently focused on data lakehouse targets such as Apache Iceberg. | - Requires Kafka operational expertise (Java, JVM tuning, distributed systems understanding). |
| - Smaller community and fewer references compared to Debezium. | - Possible latency at the Kafka storage stage; performance degradation during large-scale snapshots. | |
| - Complex connector configuration and Schema Registry management. |
When to Choose Which?
- OLake — Best when the primary goal is to quickly move operational DB data to an S3/Iceberg lakehouse for analytics; when there is insufficient staff or time to build and operate Kafka infrastructure; or when initial full load performance is critical.
- Debezium — Best when event delivery between microservices (MSA) or real-time search indexing is required; when a stable Kafka cluster is already in operation within the organization; or when a general-purpose CDC connecting diverse sources and destinations is needed.
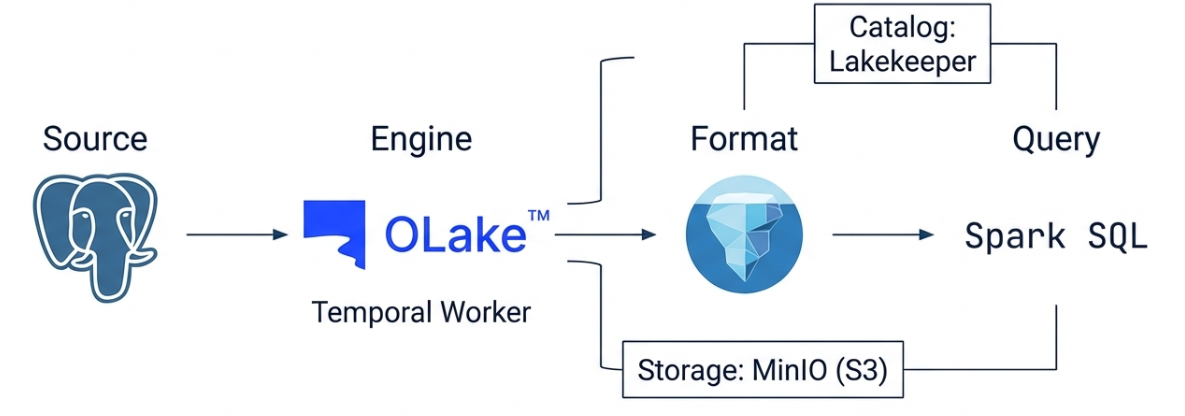
2. OLake Architecture
PostgreSQL (Source)
│ WAL (Logical Replication)
▼
OLake Worker (CDC Connector)
│ olakego/source-postgres → olake-iceberg-java-writer
▼
Lakekeeper (REST Catalog)
│
▼
Iceberg (MinIO, S3-compatible Storage)
OLake runs on top of a Temporal-based workflow engine. It reads the WAL (Write-Ahead Log) from PostgreSQL and converts it into Iceberg tables, using Lakekeeper as the Iceberg REST Catalog and MinIO as the object storage.
3. Infrastructure Setup
All services are deployed as Docker containers and connected via the olake-network bridge network.
Service Configuration
| Service | Container Name | Port | Role |
|---|---|---|---|
| OLake UI | olake-ui |
8000 | Web UI for managing CDC pipelines |
| OLake Worker | olake-temporal-worker |
— | CDC sync execution engine |
| Temporal | temporal |
— | Workflow orchestration |
| Temporal PostgreSQL | temporal-postgresql |
— | Storage for Temporal and OLake metadata |
| Source PostgreSQL | source-postgresql |
5432 | CDC source database |
| MinIO | minio |
9000 / 9001 | S3-compatible object storage |
| Lakekeeper | lakekeeper |
8181 | Iceberg REST Catalog |
Installing OLake
OLake is installed using the official one-liner script. Temporal, Worker, UI, PostgreSQL, and Elasticsearch are all deployed together.
curl -sfL https://olake.dev/install | sh
Installing Source PostgreSQL and Configuring CDC
Create a separate PostgreSQL container for the source database, independent of OLake.
docker run -d \
--name source-postgresql \
-e POSTGRES_USER=admin \
-e POSTGRES_PASSWORD=password \
-e POSTGRES_DB=mydb \
-p 5432:5432 \
postgres:16
Enable Logical Replication for CDC.
-- Modify postgresql.conf and restart
ALTER SYSTEM SET wal_level = 'logical';
-- Create a Replication Slot and Publication
SELECT pg_create_logical_replication_slot('olake_slot', 'pgoutput');
CREATE PUBLICATION olake_pub FOR ALL TABLES;
Installing MinIO and Creating a Bucket
docker run -d \
--name minio \
-e MINIO_ROOT_USER=admin \
-e MINIO_ROOT_PASSWORD=password \
-p 9000:9000 -p 9001:9001 \
minio/minio server /data --console-address ":9001"
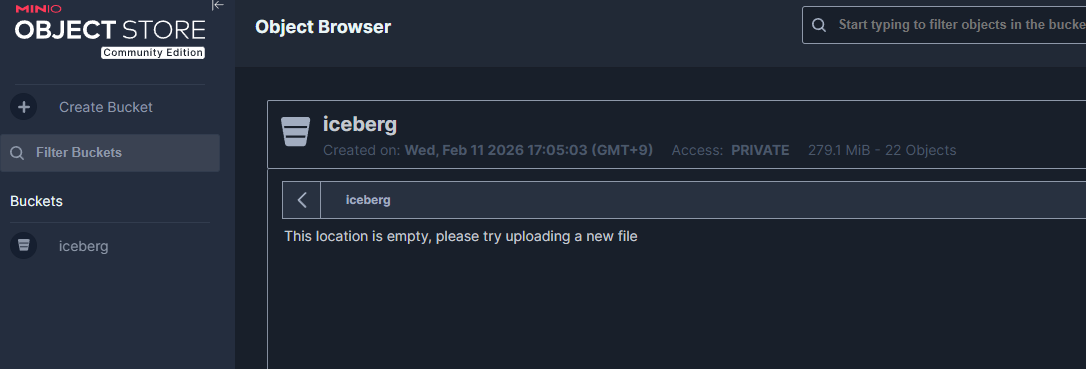
Create an iceberg bucket from the MinIO Console (http://localhost:9001).
Installing Lakekeeper
Lakekeeper is an Iceberg REST Catalog implementation that uses MinIO as its storage backend.
docker run -d \
--name lakekeeper \
-p 8181:8181 \
-e LAKEKEEPER__LISTEN_PORT=8181 \
-e LAKEKEEPER__BASE_URI=http://host.docker.internal:8181 \
lakekeeper/catalog:latest
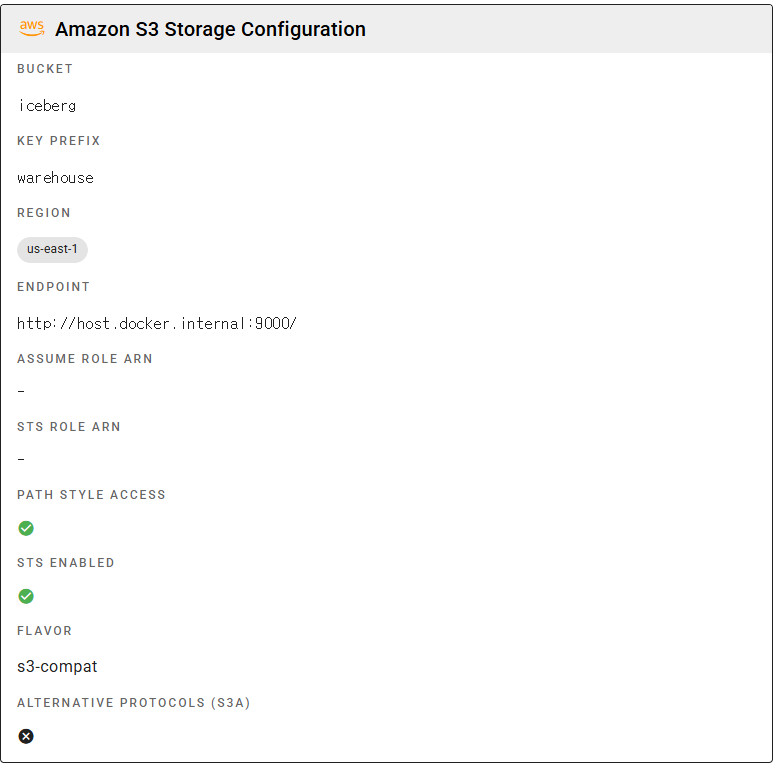
After installation, create the iceberg warehouse. This can also be done from the Lakekeeper UI (http://localhost:8181).
# Create a warehouse (S3 path: s3://iceberg/warehouse/)
curl -X POST http://localhost:8181/management/v1/warehouse \
-H "Content-Type: application/json" \
-d '{
"warehouse-name": "iceberg",
"storage-profile": {
"type": "s3",
"bucket": "iceberg",
"endpoint": "http://host.docker.internal:9000",
"path-style-access": true,
"region": "us-east-1",
"sts-enabled": true
},
"storage-credential": {
"type": "s3",
"credential-type": "access-key",
"aws-access-key-id": "admin",
"aws-secret-access-key": "password"
}
}'
Connecting to the Docker Network
Connect all services to olake-network to enable inter-container communication.
docker network connect olake-network source-postgresql
docker network connect olake-network minio
docker network connect olake-network lakekeeper
4. Loading Test Data into source_postgresql
Three datasets are loaded into the Source PostgreSQL to test the CDC pipeline.
Dataset Summary
| Table | Row Count | Description | Primary Key |
|---|---|---|---|
card_subway_month |
36,406 rows | Seoul subway monthly boarding/alighting data | line_name, station_name, use_date |
who_covid19 |
514,560 rows | WHO COVID-19 daily country-level data | date_reported, country_code |
orders |
10,000,000 rows | Test order data | order_id |
Loading Subway Data
import pandas as pd
from sqlalchemy import create_engine
engine = create_engine('postgresql://admin:password@localhost:5432/mydb')
for file in ['CARD_SUBWAY_MONTH_202501.csv', 'CARD_SUBWAY_MONTH_202502.csv']:
df = pd.read_csv(file, encoding='euc-kr')
# Rename Korean column names to English (for Iceberg compatibility)
df.columns = ['use_date', 'line_name', 'station_name',
'ride_count', 'alight_count']
df.to_sql('card_subway_month', engine, if_exists='append', index=False)
Note: OLake's Iceberg Writer replaces non-ASCII column names with
_, so using Korean column names will cause all columns to be renamed to______, resulting in a schema conflict. Always use English column names.
Loading WHO COVID-19 Data
Load 510,000 rows in chunks of 100,000.
for chunk in pd.read_csv('WHO-COVID-19-global-daily-data.csv', chunksize=100_000):
chunk.to_sql('who_covid19', engine, if_exists='append', index=False)
Loading Orders Test Data
Load 10 million order rows in chunks of 100,000.
for chunk in pd.read_csv('orders_10M.csv', chunksize=100_000):
chunk.to_sql('orders', engine, if_exists='append', index=False)
5. Configuring the OLake CDC Pipeline
Configure the Source, Destination, and Job from the OLake UI (http://localhost:8000).
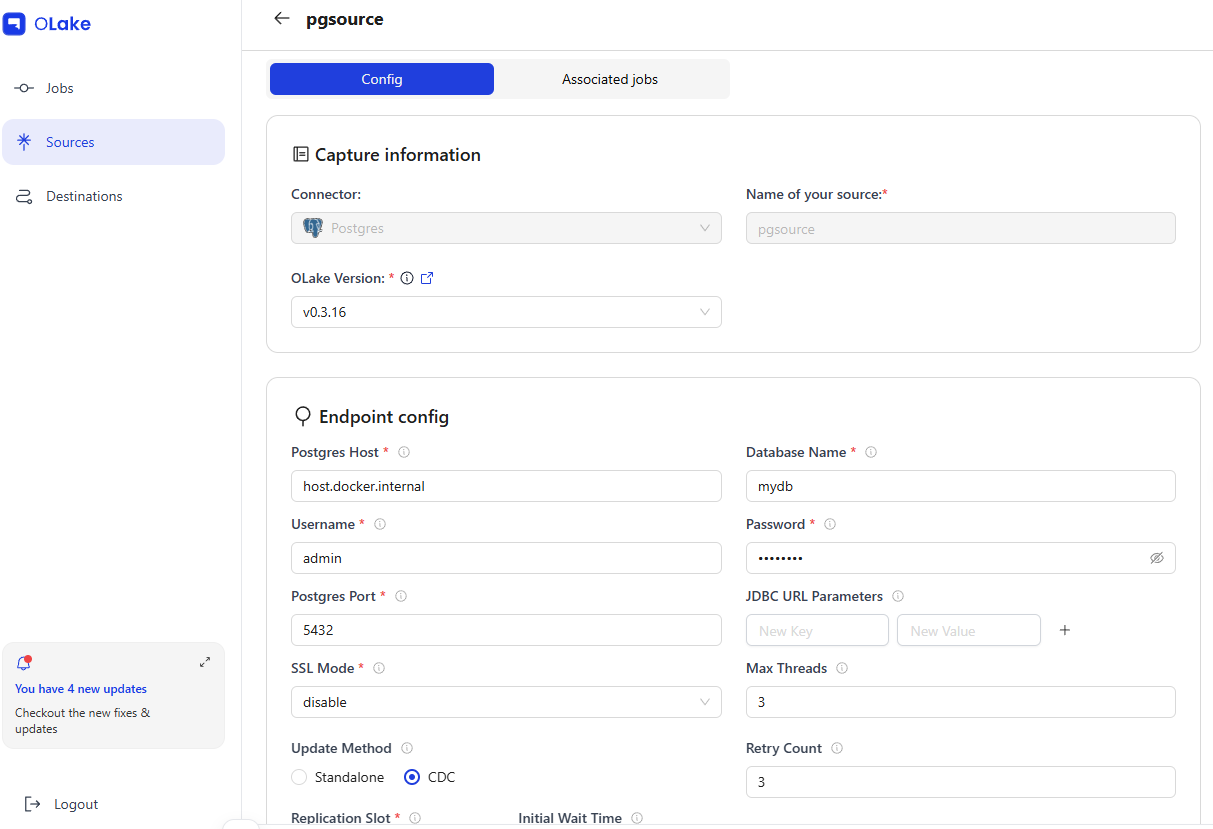
5-1. Registering a Source
| Field | Value |
|---|---|
| Name | pgsource |
| Type | postgres |
| Host | host.docker.internal:5432 |
| Database | mydb |
| Update Method | CDC (olake_pub / olake_slot) |
When a Source is registered in the OLake UI, it uses the olakego/source-postgres:v0.3.16 image to run a connection test (CHECK) and schema discovery (DISCOVER).
5-2. Registering a Destination
| Field | Value |
|---|---|
| Name | target_iceberg |
| Type | iceberg (REST Catalog) |
| REST Catalog URL | http://host.docker.internal:8181/catalog |
| S3 Endpoint | http://host.docker.internal:9000 |
| Catalog Name | olake_iceberg |
| S3 Path | iceberg |
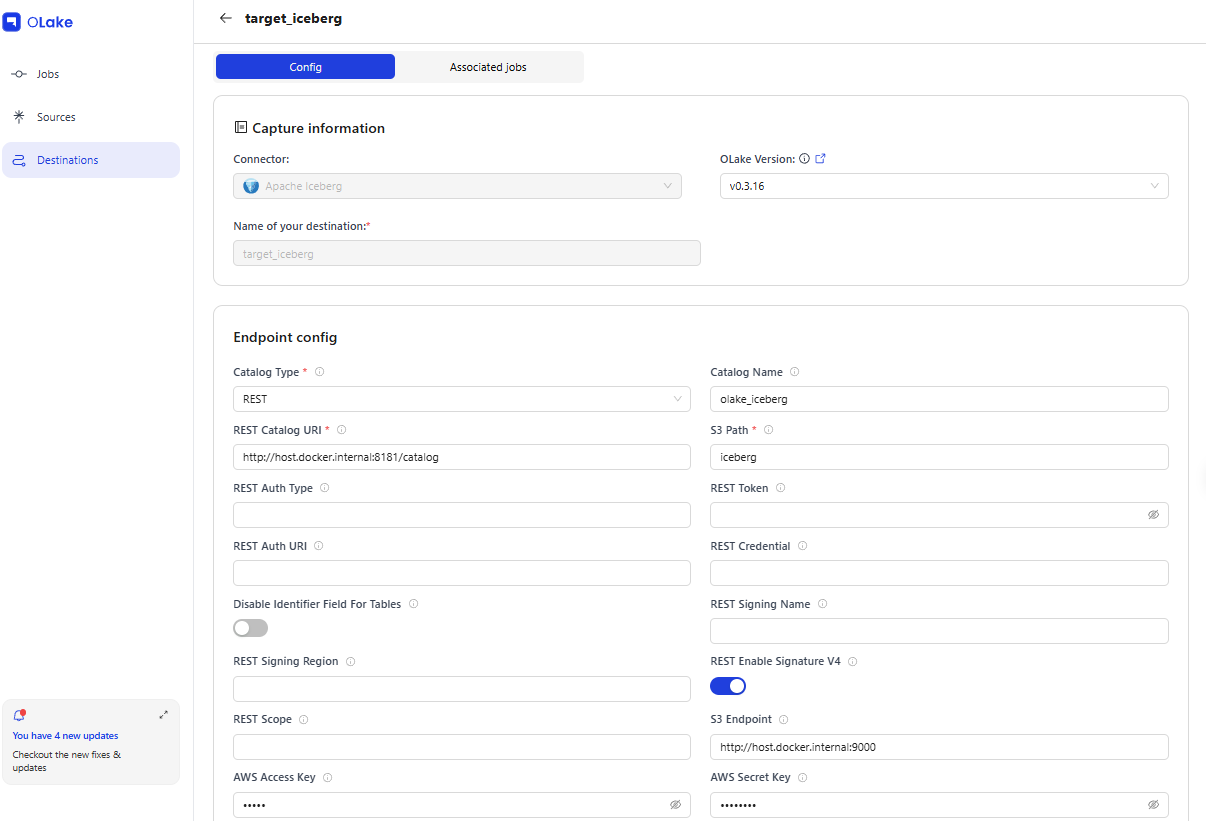
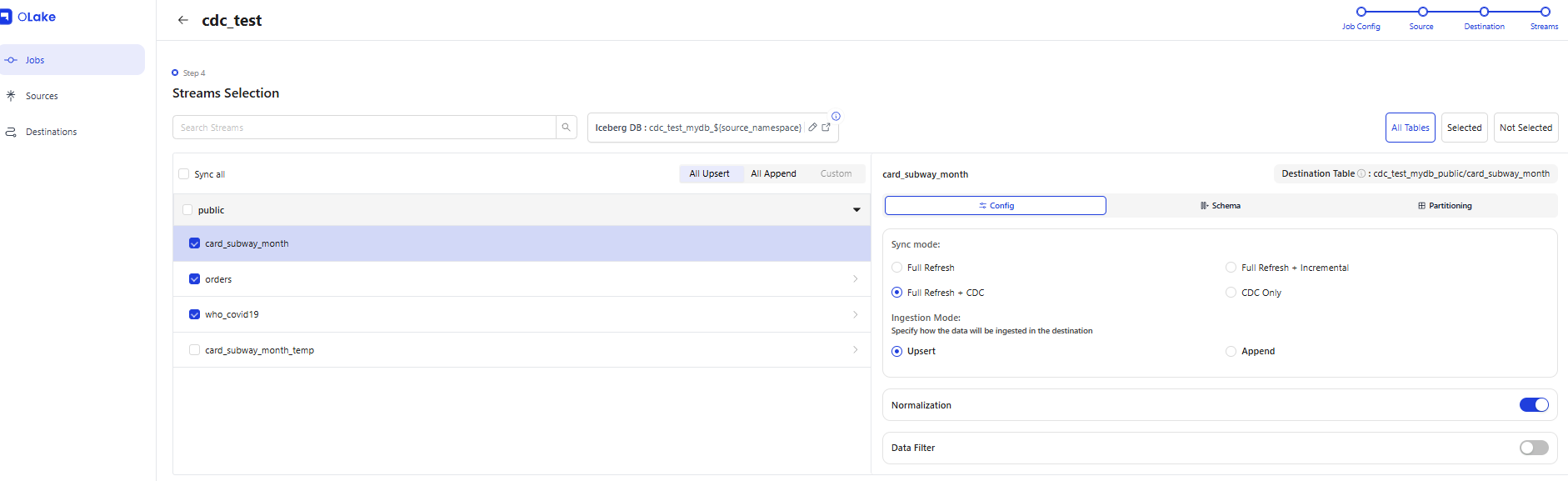
5-3. Creating a CDC Job
Link the Source and Destination, select the streams (tables) to synchronize, and create the Job. Each table's Ingestion Mode (Upsert/Append) can be selected individually. OLake also includes a Data Filter feature.
| Field | Value |
|---|---|
| Job Name | cdc_test |
| Source | pgsource |
| Destination | target_iceberg |
| Schedule | * * * * * (every minute) |
| Sync Mode | Full Refresh (initial load) + CDC |
| Ingestion Mode | Upsert |
Selected Streams
| Stream | Primary Key | Iceberg Table |
|---|---|---|
public.orders |
order_id |
cdc_test_mydb:public.orders |
public.card_subway_month |
line_name, station_name, use_date |
cdc_test_mydb:public.card_subway_month |
public.who_covid19 |
date_reported, country_code |
cdc_test_mydb:public.who_covid19 |
Once the Job is created, the OLake Worker executes a CDC sync every minute via a Temporal workflow. Each sync spawns a Docker container (olakego/source-postgres) to read the PostgreSQL WAL and writes to the Iceberg table via the Iceberg Java Writer (olake-iceberg-java-writer.jar).
How CDC Works
1. OLake Worker → Triggers a Temporal Workflow (cron every minute)
2. Docker container created (olakego/source-postgres:v0.3.16)
3. Reads WAL from the PostgreSQL Logical Replication Slot (olake_slot)
4. Delivers change records to the Iceberg Java Writer via gRPC
5. Iceberg Writer → Commits tables via the Lakekeeper REST Catalog API
6. Data files → Saved as Parquet to MinIO S3
7. State (LSN) saved → Resumes from this point in the next sync
CDC state is managed based on PostgreSQL LSN (Log Sequence Number), which tracks synchronization progress.
{
"type": "STREAM",
"global": {
"state": { "lsn": "0/9AE53D30" },
"streams": ["public.who_covid19", "public.orders", "public.card_subway_month"]
}
}
6. Querying Iceberg Tables
Set up a Spark + Jupyter environment using Docker Compose.
# docker-compose.spark.yml
services:
spark-iceberg:
image: tabulario/spark-iceberg
ports:
- "8888:8888" # Jupyter
- "8080:8080" # Spark UI
environment:
AWS_ACCESS_KEY_ID: admin
AWS_SECRET_ACCESS_KEY: password
AWS_REGION: us-east-1
Connect to the Lakekeeper catalog from Spark and run queries.
from pyspark.sql import SparkSession
# Stop any existing session (to avoid conflicts with the tabulario image's demo session)
existing = SparkSession.getActiveSession()
if existing:
existing.stop()
spark = SparkSession.builder \
.appName("iceberg-reader") \
.config("spark.sql.defaultCatalog", "lakekeeper") \
.config("spark.sql.catalog.lakekeeper", "org.apache.iceberg.spark.SparkCatalog") \
.config("spark.sql.catalog.lakekeeper.type", "rest") \
.config("spark.sql.catalog.lakekeeper.uri", "http://host.docker.internal:8181/catalog") \
.config("spark.sql.catalog.lakekeeper.warehouse", "iceberg") \
.config("spark.sql.catalog.lakekeeper.io-impl", "org.apache.iceberg.aws.s3.S3FileIO") \
.config("spark.sql.catalog.lakekeeper.s3.endpoint", "http://host.docker.internal:9000") \
.config("spark.sql.catalog.lakekeeper.s3.access-key-id", "admin") \
.config("spark.sql.catalog.lakekeeper.s3.secret-access-key", "password") \
.config("spark.sql.catalog.lakekeeper.s3.path-style-access", "true") \
.getOrCreate()
# Query data synchronized via CDC
table_name = "orders"
df = spark.table(f"""lakekeeper.cdc_test_mydb_public.{table_name}""")
total = df.count()
print(f"✓ Total: {total:,} records")
print()
df.printSchema()
df.show(10, truncate=False)
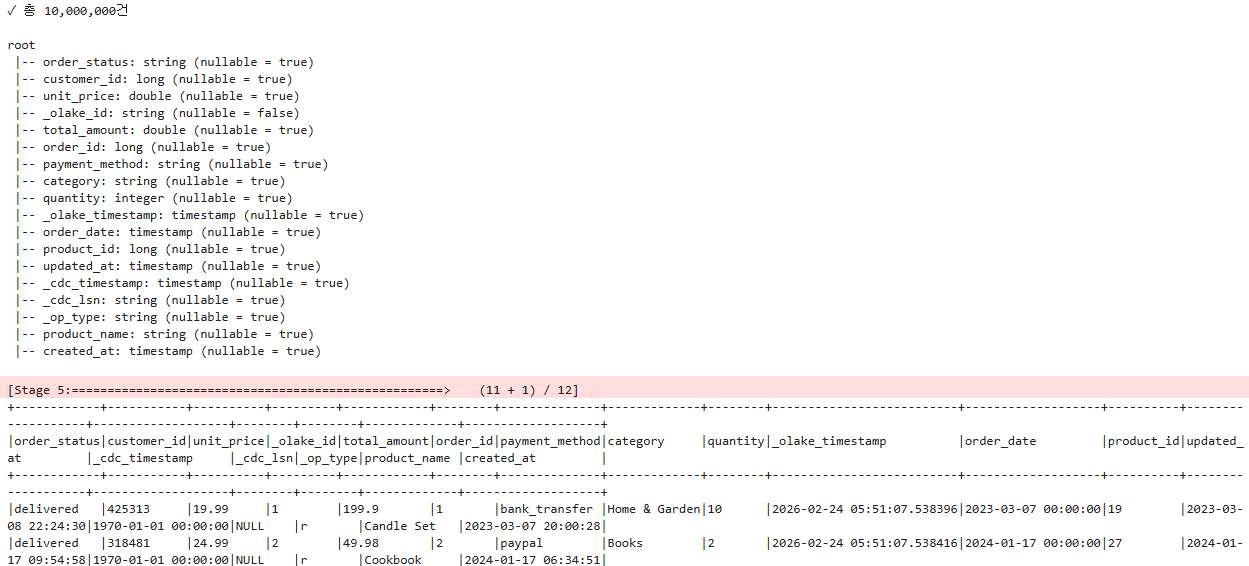
7. OLake CDC Metadata Structure
OLake stores metadata in the postgres database inside the temporal-postgresql container.
| Table | Role |
|---|---|
olake-dev-source |
Source connection information (host, DB, CDC configuration) |
olake-dev-destination |
Destination connection information (Catalog URL, S3 path) |
olake-dev-job |
Job definition (stream selection, schedule, CDC state) |
olake-dev-catalog |
Available source/destination types |
The lsn value in the CDC state represents the position in the PostgreSQL WAL. OLake updates the job state with this value after each sync completes. In subsequent syncs, only changes from this LSN onward are read and processed, implementing incremental synchronization.
8. CDC Columns in Iceberg Tables
Iceberg tables created by OLake include CDC metadata columns in addition to the original data columns.
| Column | Type | Description |
|---|---|---|
_cdc_timestamp |
timestamptz |
Timestamp of the change event in the source DB |
_olake_timestamp |
timestamptz |
Timestamp when OLake processed the record |
_op_type |
string |
Change type: r (read/initial), c (insert), u (update), d (delete) |
_olake_id |
string |
OLake internal unique ID (equality delete key) |
DELETE operations are recorded as equality delete files, with the deletion target identified by _olake_id. To exclude deleted records in Spark SQL, use the condition WHERE _op_type != 'd'.
9. Test Results
Performance
-
Initial Full Load: Even in a resource-constrained Windows environment, the initial synchronization of over 10 million records from PostgreSQL to Iceberg completed in just 162 seconds, demonstrating strong performance.
10,552,212 Records / 162 Seconds = 65,137 RPS (Records Per Second)
Job log — sync start
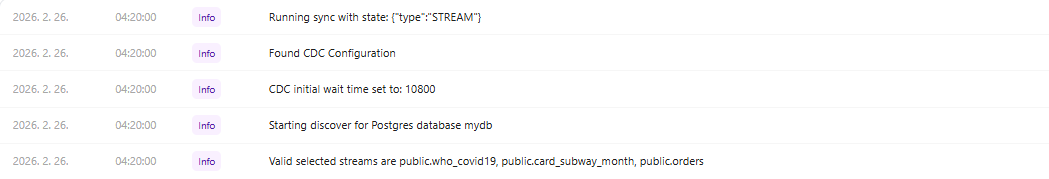
Job log — sync completed
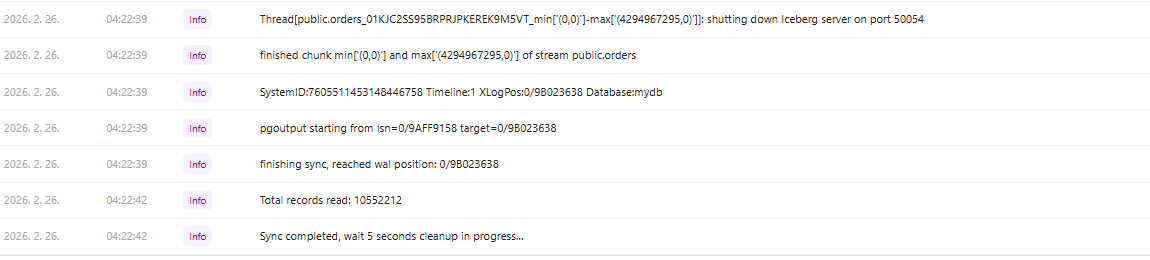
-
Incremental Sync (CDC): With a sync interval of one minute, syncs completed quickly when there were no WAL changes, and resource consumption was low since only the changed data was processed.
-
Large-Scale Data Processing: Tables of 510,000 rows (
who_covid19) and 10 million rows (orders) both maintained stable CDC synchronization after the initial load. -
Sync Container Overhead: Since a Docker container is created and destroyed for each sync, container creation costs occur even when there are no changes. For large-scale production use, adjusting the sync interval will be necessary.
Ease of Use
- Simplified Configuration: The CDC pipeline can be set up in just three steps — Source → Destination → Job — from the OLake UI, which significantly lowers the barrier to entry compared to Debezium's Kafka Connect JSON configuration.
- Built-in Connection Testing: Connection tests can be run directly from the UI when registering a Source or Destination, allowing configuration errors to be caught early.
- Automatic Stream Discovery: The Discover feature automatically retrieves the list of tables from PostgreSQL for selection, eliminating the need to manually enter table information.
Stability
- Non-ASCII Column Name Support: Non-ASCII column names (e.g., Korean) are replaced with
_by the Iceberg Writer, causing schema conflicts. Column names in the source DB must be converted to English in advance. - Long-Term Operational Stability: After running CDC continuously for several days (4 days), the
card_subway_monthstream completed over 535 syncs stably.
10. Conclusion
With OLake, a CDC pipeline from PostgreSQL to Apache Iceberg can be configured without writing any code. By registering a Source, Destination, and Job in the OLake UI, the Temporal workflow automatically handles periodic synchronization.
| Component | Choice | Role |
|---|---|---|
| CDC Source | PostgreSQL + Logical Replication | WAL-based change capture |
| CDC Engine | OLake (Temporal Worker) | Change data extraction and delivery |
| Table Format | Apache Iceberg | ACID transactions, schema evolution, time travel |
| REST Catalog | Lakekeeper | Iceberg table metadata management |
| Object Storage | MinIO | Parquet data file storage |
| Query Engine | Spark SQL | Iceberg table querying |
Compared to Kafka-based Debezium, OLake offers simpler infrastructure and easier initial setup. However, it still has areas of immaturity, including limited monitoring capabilities and lack of support for non-ASCII column names. It can be a strong choice for teams that need to quickly build a lakehouse-centric data pipeline.
